 If you wish to visualize them on Paraview with the correct physical coordinates, I think the only way is to create a xmdf file. I use them to deal with another issue actually which I am not sure if it is important for you or not, NetCDF files are saved based on array index. But anything that is opt-in, separate download etc.Xmdf files helped me to read NetCDF files with older versions of Paraview as well. If one manage something like -DHDF5_BUILD_PLUGINS=YES, where “YES” is the default, in the main HDF5 cmake build system, then it might work out. Gzip compression currently works because everyone has it in the default HDF5 build, but that is so incredibly slow that it is practically useless for any purpose. I would need to distribute plugins for Mac, Linux and Windows, which I have no possibility to do, I do not even have a Windows or Mac computer to compile such a thing on… And even that is impossible, because 80% of the users would still not be able to get the plugin running properly. Also support for a number of file formats. So If I compress my dataset with Zstd on the CFD code side, I will need to distribute the plugin along with the dataset to everyone that want to have a look, because the Paraview installation does not have the plugin installed. No native support for HDF5, however, ParaView supports a container format XDMF which uses HDF5 for actual data. Paraview ships its own HDF5 that might be a complete different version, built with different compilers and with different features. But then, when I go to a random system for postprocesisng to open it in Paraview, we instantly get a problem. I can include as many plugins and filters as I wish in my CFD code. The systems used for processing and visualization are completely “random”, could be any system really. The data is typically computed on a HPC system, and the visualization happens somewhere else. The files can be opened with tools like Paraview, and we also use a lot of custom Python programs for processing the data. The problems with filters as separate plugins is the following: Having official in-library Zstd compression would be awesome and allow using compression in many new cases! Gzip is too slow for practical use, and if I use Zstd nobody will be able to open my files, because they lack the plugin. Paraview binaries ship with HDF5 support, then my files are readable. I do for instance rely on HDF5 for storing simulation data, and this data is opened with Paraview. If I suddenly start using filter-plugins, that is no longer the case. When only using official library features I can assume that the HDF5 files I make is universally readable everywhere. 
In my opinion, one of the great things with HDF5 is the portability. I am aware that there is a filter-plugin for it, but it’s not quite the same as built-in support.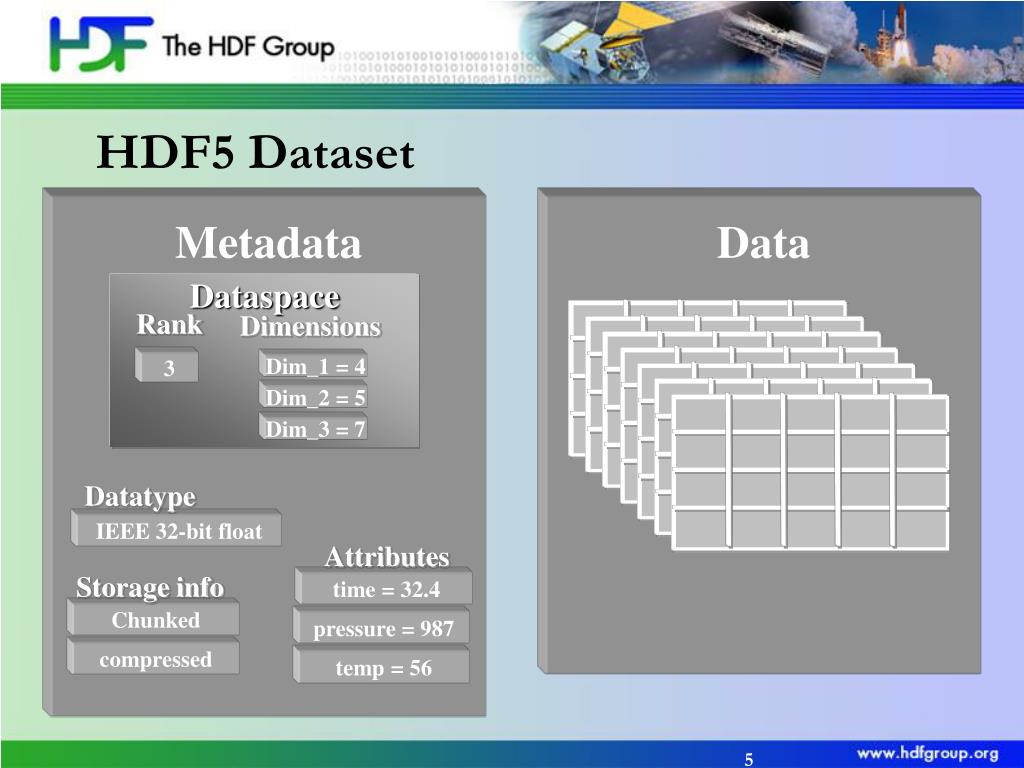 
Zstd have gained tremendous popularity in the last years, due to extremely efficient compression/decompression (thousands of MB/s on a single core) together with a quite acceptable compression ratio (on par with gzip).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
